Pygenprop’s Documentation¶
Overview¶
Pygenprop is a python library for programmatic exploration and usage of the EBI Genome Properties database.
Features¶
At its core the library contains four core components:
- An object model for representing the Genome Properties database as a rooted direct acyclic graph.
- A parser for Genome Properties database flat files.
- A parser for Genome Properties assignment longform files.
- A parser for InterProScan TSV files.
Implementation¶
In-memory Data Structures¶
Pygenprop generates a series of in-memory data structures representing the Genome Properties database. A series of steps support the existence of each genome property, and pieces of evidence support the existence of each of these steps. Steps are assigned as YES or NO based on evidence, such as the presence of InterPro matches or support for the existence of child properties. Some steps can be required whereas others can be optional. If all required steps are assigned YES, then the genome property is also assigned YES. If only a subset of the required steps are assigned YES, then the property may be assigned PARTIAL or NO. Because some properties rely on the assignments of others, the Genome Properties database forms a rooted directed acyclic graph (DAG). We refer to this data structure as a “property tree”, although child properties can have multiple parent properties.
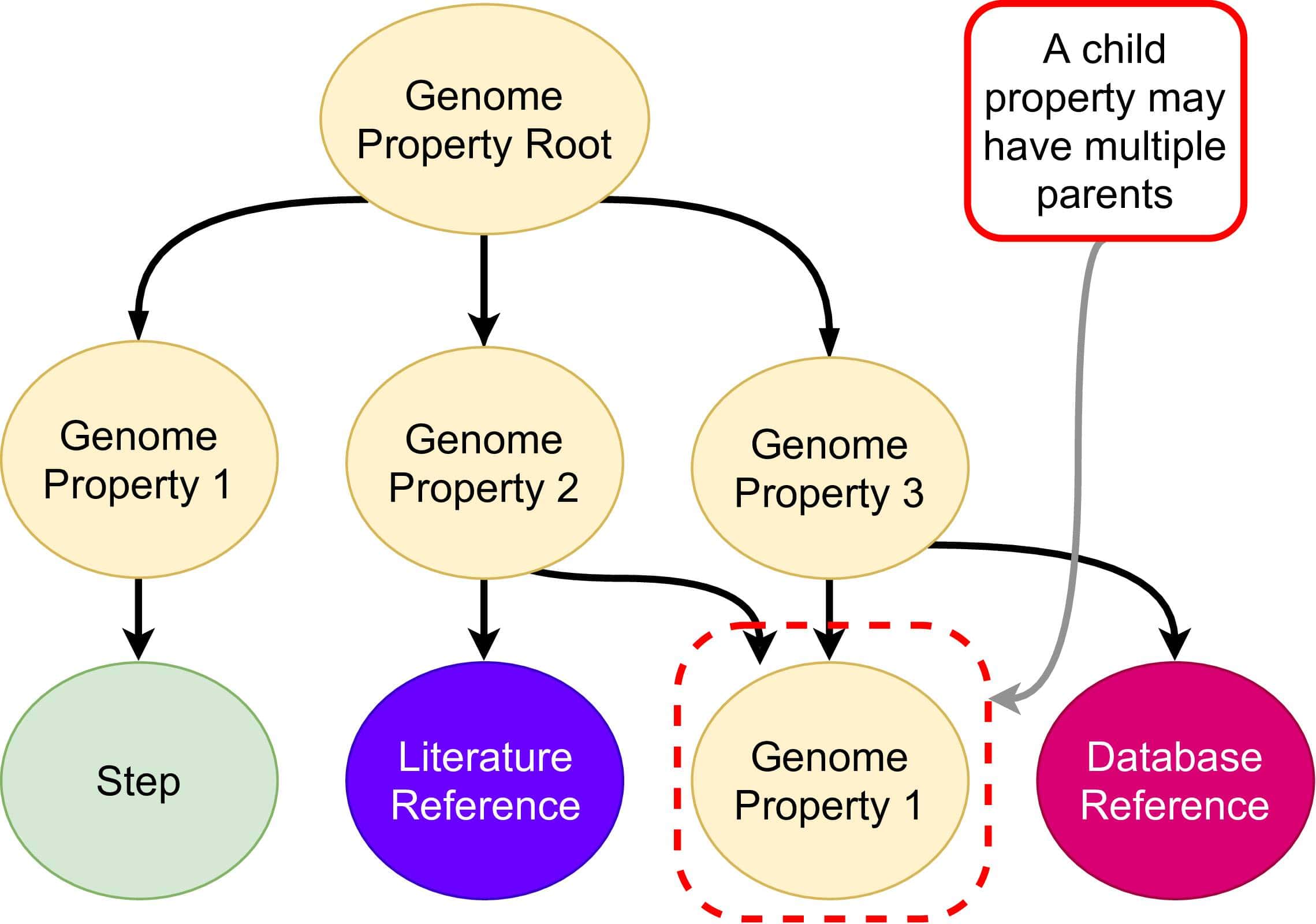
Some property objects are the children of others. Database reference, literature reference and step objects are children of property objects.
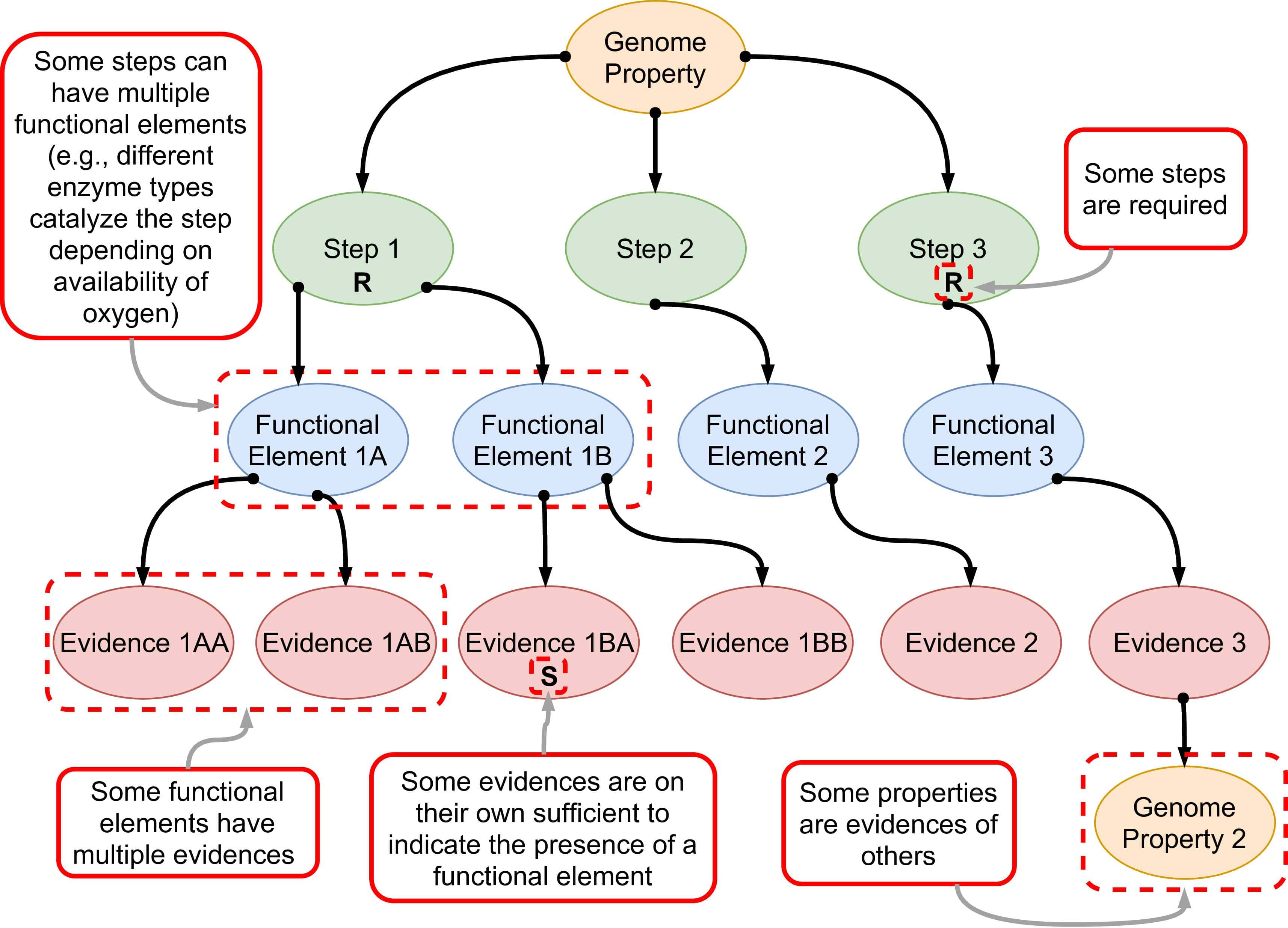
Each property is supported by step, functional element, and evidence objects.
Pygenprop loads the entire Genome Properties database into main memory and represents the data using simple Python data structures. A custom parser for Genome Properties database flat files instantiates data classes into objects. The core object is the property tree (a doubly linked rooted DAG) with one or more child property objects. Each property links with both its parent and child, allowing for climbing up and down the DAG structure. Properties are also indexed by unique identifiers that are stored within a dictionary, which allows for rapid queries. The property tree class has shortcuts for accessing the root genome property (GenProp0065), leaf genome properties, and any property by its unique identifier. Each genome property also includes child objects containing additional information, including external database references (to the equivalent pathway in KEGG, MetaCyc , and IUBMB Enzyme Reactions, literature supporting the existence of the property on PubMed, steps, functional elements, and step evidence. Functional element objects are children of steps, and step evidence objects are children of functional elements. The functional element is a new data class that Pygenprop adds to Genome Properties at runtime. The parser places functional elements in between steps and evidence to accommodate cases in which a reaction may be catalyzed by multiple disjoint enzyme classes. For example, step #1 of GenProp2023 (caspase activation) can be catalyzed either by caspase-2 or granzyme B (see Github Issue ). We represent these enzymes as two separate functional elements with their respective evidence.
Property Assignment¶
We have replicated the code found in the Genome Properties Perl library, which calculates property assignments from InterProScan annotation tab-separated values (TSV) files. As with the Genome Properties Perl library, InterPro matches for an organism’s genome are parsed and extracted from the TSV file, then dereplicated into a unique set. Subsequently, Pygenprop recursively calculates, from leaf to root, YES, NO, or PARTIAL property assignments depending on the presence or absence of signatures in the above set and a threshold for the minimum number of required steps needed for a PARTIAL assignment of each property. Pygenprop contains parsers for Genome Properties assignment files (created by the original Perl library) and InterProScan TSV files.
Assignment Results¶
We include a results class for containing the full property set and step assignments across multiple organisms. Once instantiated, objects of this class contain two pandas DataFrame objects, one for property assignments and one for step assignments. Each object comprises rows of assignments by columns of samples. The results class also contains methods for filtering these DataFrames down to assignments for properties shared between organisms or assignments for specific properties of interest.
Integration with other frameworks¶
A key advantage of Pygenprop is its ability to be integrated within existing Python-based bioinformatics and data science frameworks. Pygenprop can be imported into Jupyter Notebooks and used in interactive analyses. Property assignments generated by Pygenprop are stored in pandas DataFrames , allowing for filtering (e.g., finding properties that differ between organisms) or clustering (e.g., using scikit-learn ). The DataFrames are also integrated readily with machine learning libraries, such as scikit-learn, Pytorch , or TensorFlow, which can drive downstream applications such as phenotype classification. Calculations performed using pandas can be visualized using Python libraries, such as Seaborn and Bokeh. Pygenprop can be used with Python-based bioinformatics web applications that require parsing and assignment of Genome Properties data. Finally, Pygenprop can be used to compute genome property assignments from a simple list of InterPro signatures, such as those imported from remote web application programming interfaces (APIs).
Tutorial¶
A basic tutorial can be found on our Github README. More detailed documentation on each module’s API can be found in the Module Index.